 Ubiquitin (Ub) is a small protein that is – you guessed it – ubiquitous in eukaryotes, where it is highly conserved with only 3 differences between the Ub of yeast and humans. (It is found in neither eubacteria nor archaebacteria.) The translation products of Ub genes are fusion proteins, and Ub genes can exist in two states (a) fused to a ribosomal protein gene to yield a Ub-ribosomal fusion protein, and (b) linear repeats that yield a translation product that comprises a linear chain of Ub-molecules (a polyubiquitin molecule). Following synthesis of the fusion proteins, Ub-C-term hydrolase cleaves the fusion proteins at the C-terminal end of Ub. Hydrolysis either liberates an individual Ub and ribosomal protein or it liberates a set of Ub monomers from the polyubiquitin.
Ubiquitin (Ub) is a small protein that is – you guessed it – ubiquitous in eukaryotes, where it is highly conserved with only 3 differences between the Ub of yeast and humans. (It is found in neither eubacteria nor archaebacteria.) The translation products of Ub genes are fusion proteins, and Ub genes can exist in two states (a) fused to a ribosomal protein gene to yield a Ub-ribosomal fusion protein, and (b) linear repeats that yield a translation product that comprises a linear chain of Ub-molecules (a polyubiquitin molecule). Following synthesis of the fusion proteins, Ub-C-term hydrolase cleaves the fusion proteins at the C-terminal end of Ub. Hydrolysis either liberates an individual Ub and ribosomal protein or it liberates a set of Ub monomers from the polyubiquitin.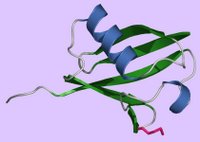 (left - Ubiquitin ribbon - click to enlarge - molecular surface ubiquitin - above, right ) Ubiquitin is a 76-aa heat-stable protein that folds into a compact globular structure. Ub may exist as a free monomer in eukaryotic cells, or it may be co-valently attached to itself (polyubiquitin) or other proteins (conjugated Ub). Single Ub molecules or Ub chains conjugate (ATP-dependent) to proteins through a covalent bond between the Ub's C-terminal glycine (pink) and the protein's lysine side chains.
(left - Ubiquitin ribbon - click to enlarge - molecular surface ubiquitin - above, right ) Ubiquitin is a 76-aa heat-stable protein that folds into a compact globular structure. Ub may exist as a free monomer in eukaryotic cells, or it may be co-valently attached to itself (polyubiquitin) or other proteins (conjugated Ub). Single Ub molecules or Ub chains conjugate (ATP-dependent) to proteins through a covalent bond between the Ub's C-terminal glycine (pink) and the protein's lysine side chains.The Ubiquitin Proteasome Pathway []UPP[] is central to the regulation of almost all cellular processes including:
● Antigen processing
● Apoptosis
● Biogenesis of organelles and ribosomes
● Cell cycle and division
● DNA transcription and DNA repair
● Cellular differentiation and development
● Immune response and inflammatory response
● Neural and muscular degeneration
● Morphogenesis of neural networks
● Modulation of cell surface receptors, ion channels and the secretory pathway
● Response to cellular stress and extracellular signaling molecules
● Viral infection
Ub regulates cellular protein turnover (protein degradation) by closely regulating the degradation of specific proteins by cellular machinery, enabling cells to rapidly eliminate other regulatory proteins such as transcription factors. Proteins destined* for degradation are first conjugated to Ub, assisted by Ub-activating (E1), Ub-conjugating (E2), and Ub-ligase enzymes, before the Ub-protein complex is transported to the proteasome for hydrolysis by proteases. The E1 enzymes activates the C-terminal cysteine residue of a ubiquitin monomer to a high-energy thiolester bond, which is then transferred to a reactive cysteine residue of the E2 enzyme. The E3 ligase enzyme then conducts the transfer of ubiquitin to the ε-amino group of a reactive lysine residue of the protein destined for degradation, and then the Ub-protein complex is escorted to the proteasome for hydrolysis by proteases.
[] 26s proteasome [] 3D structure of targetted peptide [] barrel and subunits of proteasome [] proteasome assembly [] proteasome engine [] substrate swapping [] Ub-proteasome pathway [] yeast 20s proteasome ribbon, yeast 20s proteasome down the barrel, surface with bound aldehyde indicators Џ animation - proteasome Џ animation - importing, unfolding, hydrolyzing Џ
Deubiquitinating enzymes remove Ub from substrate proteins, rescuing de-conjugated proteins from degradation.
: diag. Ub-proteasome pathway : diag. ubiquitin-mediated degradation : diag. UBL/UBL proteins : MPEG movies ~ Localisation during meiosis I ~ Localisation during meiosis II ~ Localisation during mitosis ~ Localisation during karyogamy ~ Localisation during "horsetail movement"
Ub conjugation ensures it a vital role in cellular process:
1. conjugated to the protein cyclin during the G1 phase of mitosis ensures an important role in regulating the cell cycle.
2. DNA repair
3. regulation of transcription
4. embryogenesis
5. apoptosis
6. receptor internalisation
Examples of role of ubiquitination:
A protein that mediates cell adhesion by binding cytoskeletal F-actin, signal transduction pathways mediated by the EGF and proto-oncogenes src and Wnt-1. When the Wnt signaling pathway is inactive, the cytoplamsic pool of β-catenin is degraded by a complex including casein kinase 1 (CK1), glycogen synthase kinase 3 (GSK), APC and Axin, which phosphorylate β-catenin (PPPP). The phosphorylated β-catenin is recognized by the ubiquitin ligase component β-TrCP, which together with Skp1, Cul1 and the E1 and E2 ubiquitination components, mediates the ubiquitylation of β-catenin, directing it to degradation by the 26S proteasome.)[s]
Aberrant UB-mediated protein degradation has been implicated in a number of pathological conditions, particularly neurodegenerative disorders that involve protein aggregation and inclusion body formation, where protein misfolding may play a role –Alzheimer’s disease, Parkinson’s disease, Huntington’s disease, and ALS.
* Determinants of half-life of proteins:
1. 'N-degron', N-end rule according to N-terminal amino acid (Ser long-lived → Asp short-lived) ,
2. amino acid sequences such as the PEST sequence, rich in proline, glutamic acid, serine, and threonine (if not masked by covalent attachment of phosphate groups to the side chains of certain amino acids)
3. exposure of degradation-signals in a partially unfolded state, allowing the signals to interact with the Ub machinery, causing the protein to become tagged by Ub. This reaction appears to be hindered by chaperone activity.
[more detail 1]
▲ Top ▲
No comments:
Post a Comment